%matplotlib inline
import numpy as np
import scipy as sp
import matplotlib as mpl
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import pandas as pd
pd.set_option('display.width', 500)
pd.set_option('display.max_columns', 100)
pd.set_option('display.notebook_repr_html', True)
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("poster")Identifiability in Bayesian Models
How parameter redundancy breaks MCMC convergence — and how priors can help.
We generate some test data from \(N(0,1)\):
from scipy.stats import norm
data = norm.rvs(size=100)
dataarray([-1.40705399, 0.47855778, -0.09668042, 1.73340459, -0.0935003 ,
1.51884902, 0.14321649, -0.97385946, -1.25932683, -0.11006569,
0.39172028, -0.28017972, -1.1848513 , 0.7813908 , 1.21234791,
0.58837838, 0.44120785, -1.31492176, 0.56251823, -1.17382197,
-0.5036201 , 0.66069273, 0.9170503 , 0.0818321 , -0.39673409,
-0.17703474, 0.622938 , -0.31216903, -0.76925065, -1.27985395,
0.25207792, -0.71459936, -1.5035201 , -0.20299391, 1.9505443 ,
1.82787502, 0.10564024, -0.65272199, 0.11243971, 1.32859278,
1.07419161, 1.54344363, 0.47665397, 0.83532311, -1.16837571,
-0.66616421, -1.5110644 , -0.88473802, -1.30381742, 1.09177324,
-0.34623827, 1.04072798, -0.40245543, -0.93406504, -0.23590713,
-0.90397347, -2.04925936, 1.19272474, 0.72976297, 0.12442893,
-1.38611462, -1.11220143, -1.41744909, -0.26507423, 0.46588182,
-1.33920462, -1.44011436, -0.62538301, -0.54746824, 2.14935439,
0.69735202, 0.85241117, 0.47595123, 0.05334232, -0.0822768 ,
-0.52292048, -0.57991741, 0.48867043, 0.85620906, -0.82954922,
1.14262662, -0.1381081 , -0.92574713, 0.59721557, -0.19176936,
0.84759029, 0.35331409, 1.58063441, -0.9334769 , -0.6522197 ,
-0.69524332, -1.30607046, -0.62902851, 1.1504571 , 0.27805355,
2.46018131, -0.20520832, 1.15109449, 0.5690628 , 0.30671181])We fit this data using the following model:
\[ y \sim N(\mu, \sigma)\\ \mu = \alpha_1 + \alpha_2\\ \alpha_1 \sim Unif(-\infty, \infty)\\ \alpha_2 \sim Unif(-\infty, \infty)\\ \sigma \sim HalfCauchy(0,1) \]
import pymc as pm
import arviz as azIn our sampler, we have chosen cores=2 which allows us to run on multiple processes, generating two separate chains.
with pm.Model() as ni:
sigma = pm.HalfCauchy("sigma", beta=1)
alpha1=pm.Uniform('alpha1', lower=-10**6, upper=10**6)
alpha2=pm.Uniform('alpha2', lower=-10**6, upper=10**6)
mu = pm.Deterministic("mu", alpha1 + alpha2)
y = pm.Normal("data", mu=mu, sigma=sigma, observed=data)
stepper=pm.Metropolis()
traceni = pm.sample(100000, step=stepper, cores=2)Multiprocess sampling (2 chains in 2 jobs)
CompoundStep
>Metropolis: [sigma]
>Metropolis: [alpha1]
>Metropolis: [alpha2]/Users/rahul/Library/Caches/uv/archive-v0/-9zMAs8POxRAxupzaV9ft/lib/python3.14/site-packages/rich/live.py:260:
UserWarning: install "ipywidgets" for Jupyter support
warnings.warn('install "ipywidgets" for Jupyter support')
Sampling 2 chains for 1_000 tune and 100_000 draw iterations (2_000 + 200_000 draws total) took 12 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for detailspm.model_to_graphviz(ni)az.summary(traceni)| mean | sd | hdi_3% | hdi_97% | mcse_mean | mcse_sd | ess_bulk | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|---|
| sigma | 0.979 | 0.070 | 0.850 | 1.111 | 0.000 | 0.000 | 25548.0 | 27098.0 | 1.00 |
| alpha1 | 1.426 | 5.276 | -9.282 | 8.457 | 3.263 | 1.436 | 3.0 | 26.0 | 1.73 |
| alpha2 | -1.429 | 5.275 | -8.445 | 9.305 | 3.263 | 1.436 | 3.0 | 25.0 | 1.73 |
| mu | -0.003 | 0.098 | -0.181 | 0.186 | 0.001 | 0.000 | 26626.0 | 27051.0 | 1.00 |
az.plot_trace(traceni);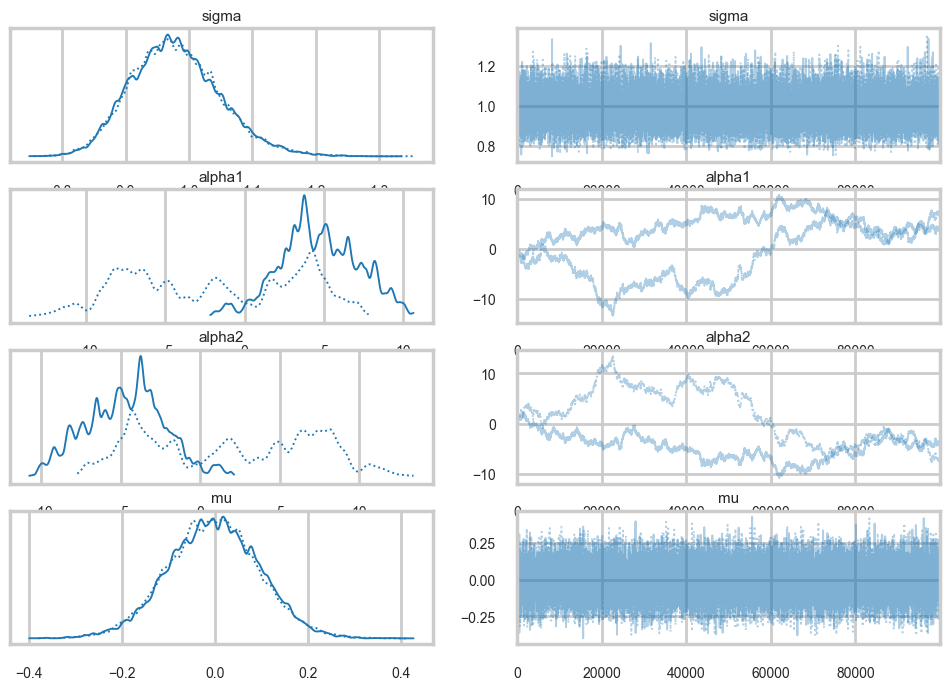
Look at our traces for \(\alpha_1\) and \(\alpha_2\). These are bad, and worse, they look entirely different for two chains. Despite this, \(\mu\) looks totally fine. Our trac
df=traceni.posterior.to_dataframe()
df.corr()| sigma | alpha1 | alpha2 | mu | |
|---|---|---|---|---|
| sigma | 1.000000 | 0.002777 | -0.002880 | -0.005532 |
| alpha1 | 0.002777 | 1.000000 | -0.999828 | 0.021611 |
| alpha2 | -0.002880 | -0.999828 | 1.000000 | -0.003077 |
| mu | -0.005532 | 0.021611 | -0.003077 | 1.000000 |
Just like in our uncentered regression example, we have \(\alpha_1\) and \(\alpha_2\) sharing information: they are totally negatively correlated and unidentifiable. Indeed our intuition probably told us as much.
az.plot_autocorr(traceni);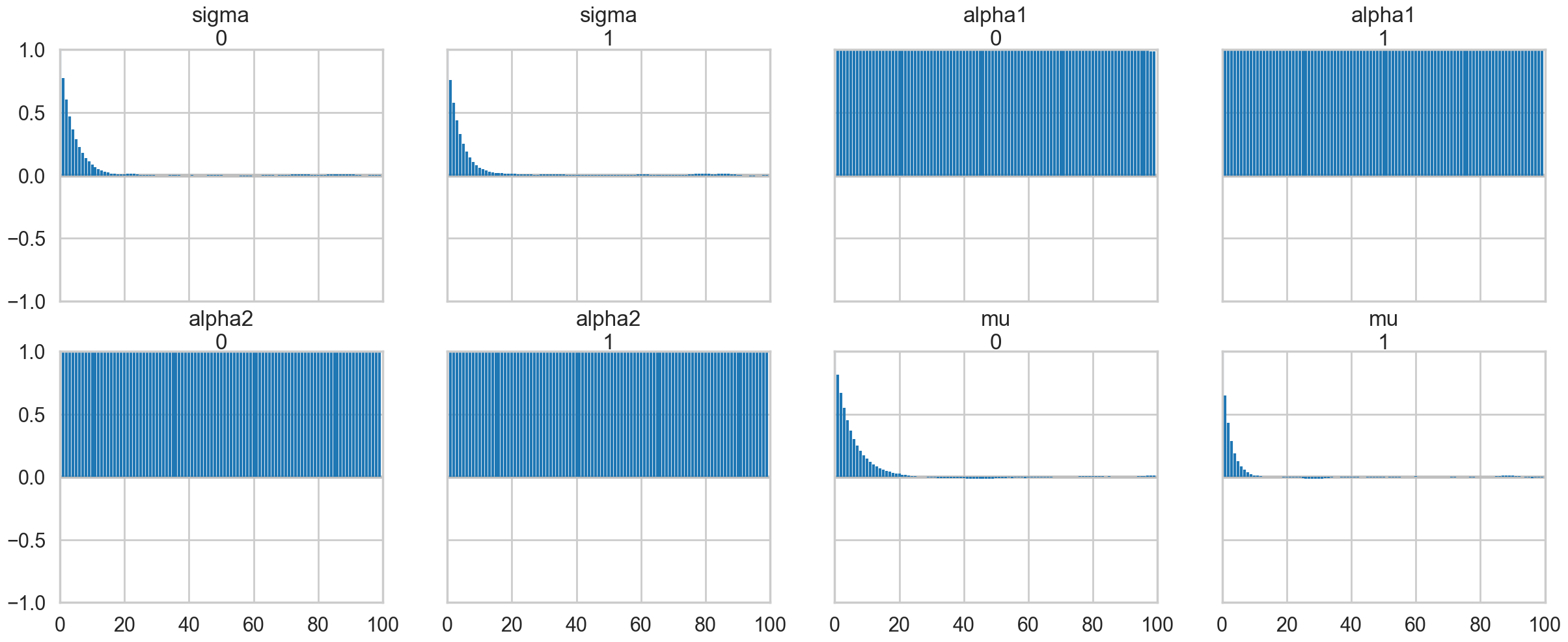
A look at the effective number of samples using two chains tells us that we have only one effective sample for \(\alpha_1\) and \(\alpha_2\).
az.ess(traceni)<xarray.Dataset> Size: 32B
Dimensions: ()
Data variables:
sigma float64 8B 2.555e+04
alpha1 float64 8B 3.074
alpha2 float64 8B 3.073
mu float64 8B 2.663e+04
Attributes:
created_at: 2026-03-06T18:44:19.320198+00:00
arviz_version: 0.23.4
inference_library: pymc
inference_library_version: 5.28.1
sampling_time: 12.227087020874023
tuning_steps: 1000The Gelman-Rubin statistic is awful for them. No convergence.
az.rhat(traceni)<xarray.Dataset> Size: 32B
Dimensions: ()
Data variables:
sigma float64 8B 1.0
alpha1 float64 8B 1.731
alpha2 float64 8B 1.732
mu float64 8B 1.0
Attributes:
created_at: 2026-03-06T18:44:19.320198+00:00
arviz_version: 0.23.4
inference_library: pymc
inference_library_version: 5.28.1
sampling_time: 12.227087020874023
tuning_steps: 1000Its going to be hard to break this unidentifiability. We try by forcing \(\alpha_2\) to be negative in our prior
with pm.Model() as ni2:
sigma = pm.HalfCauchy("sigma", beta=1)
alpha1=pm.Normal('alpha1', mu=5, sigma=1)
alpha2=pm.Normal('alpha2', mu=-5, sigma=1)
mu = pm.Deterministic("mu", alpha1 + alpha2)
y = pm.Normal("data", mu=mu, sigma=sigma, observed=data)
#stepper=pm.Metropolis()
#traceni2 = pm.sample(100000, step=stepper, cores=2)
traceni2 = pm.sample(100000)Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [sigma, alpha1, alpha2]/Users/rahul/Library/Caches/uv/archive-v0/-9zMAs8POxRAxupzaV9ft/lib/python3.14/site-packages/rich/live.py:260:
UserWarning: install "ipywidgets" for Jupyter support
warnings.warn('install "ipywidgets" for Jupyter support')
Sampling 4 chains for 1_000 tune and 100_000 draw iterations (4_000 + 400_000 draws total) took 66 seconds.pm.model_to_graphviz(ni2)Notice we are using the built in NUTS sampler. It takes longer but explores the distributions far better. This is directly related to our priors imposing regions. I could not even run the previous sampler in any reasonable time in NUTS.
az.plot_trace(traceni2);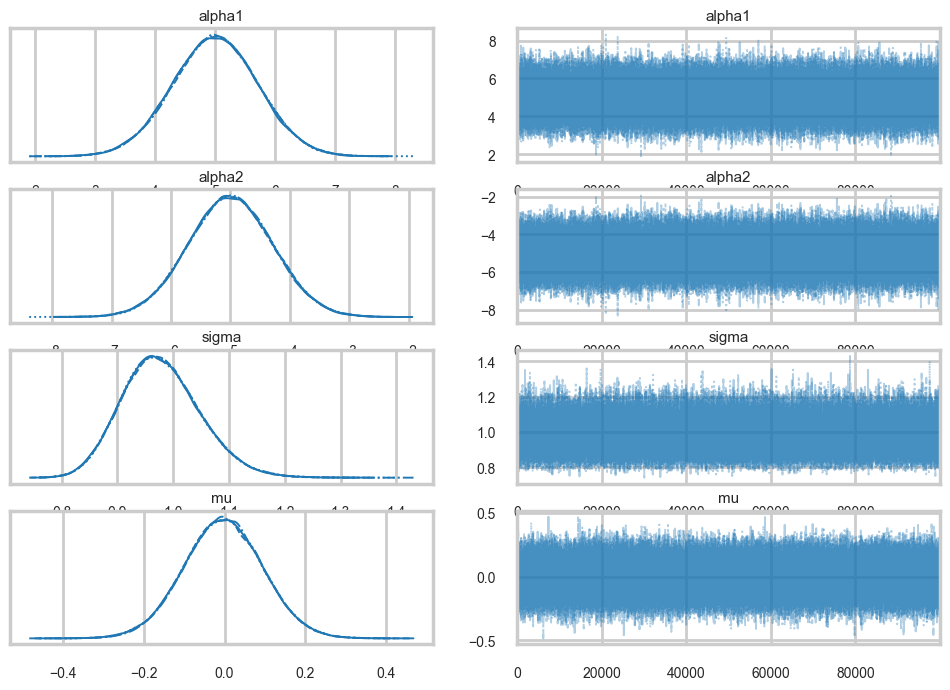
Our extremely strong priors have helped us do a much better job.
az.summary(traceni2)| mean | sd | hdi_3% | hdi_97% | mcse_mean | mcse_sd | ess_bulk | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|---|
| alpha1 | 5.000 | 0.710 | 3.679 | 6.354 | 0.002 | 0.002 | 127028.0 | 147579.0 | 1.0 |
| alpha2 | -5.003 | 0.710 | -6.331 | -3.655 | 0.002 | 0.002 | 127359.0 | 147791.0 | 1.0 |
| sigma | 0.979 | 0.070 | 0.849 | 1.111 | 0.000 | 0.000 | 176191.0 | 167933.0 | 1.0 |
| mu | -0.004 | 0.098 | -0.189 | 0.180 | 0.000 | 0.000 | 418836.0 | 308070.0 | 1.0 |
Our effective sample size is still poor and our traces still look dodgy, but things are better.
az.ess(traceni2)<xarray.Dataset> Size: 32B
Dimensions: ()
Data variables:
alpha1 float64 8B 1.27e+05
alpha2 float64 8B 1.274e+05
sigma float64 8B 1.762e+05
mu float64 8B 4.188e+05
Attributes:
created_at: 2026-03-06T18:45:36.679458+00:00
arviz_version: 0.23.4
inference_library: pymc
inference_library_version: 5.28.1
sampling_time: 66.3943829536438
tuning_steps: 1000az.rhat(traceni2)<xarray.Dataset> Size: 32B
Dimensions: ()
Data variables:
alpha1 float64 8B 1.0
alpha2 float64 8B 1.0
sigma float64 8B 1.0
mu float64 8B 1.0
Attributes:
created_at: 2026-03-06T18:45:36.679458+00:00
arviz_version: 0.23.4
inference_library: pymc
inference_library_version: 5.28.1
sampling_time: 66.3943829536438
tuning_steps: 1000..and this shows in our Gelman-Rubin statistics as well…
traceni2.posterior.to_dataframe().corr()| alpha1 | alpha2 | sigma | mu | |
|---|---|---|---|---|
| alpha1 | 1.000000 | -0.990497 | 0.000240 | 0.063937 |
| alpha2 | -0.990497 | 1.000000 | -0.000245 | 0.073925 |
| sigma | 0.000240 | -0.000245 | 1.000000 | -0.000042 |
| mu | 0.063937 | 0.073925 | -0.000042 | 1.000000 |
..but our unidentifiability is still high when we look at the correlation. This reflects the fundamental un-identifiability and sharing of information in our model since \(\mu = \alpha_1 +\alpha_2\): all the priors do is artificially peg one of the parameters. And once one is pegged the other is too because of the symmetry.